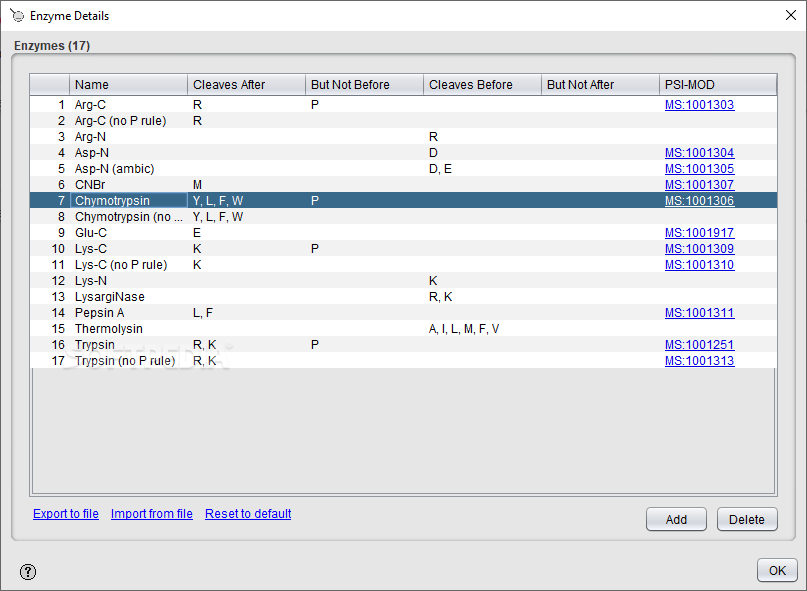
> The command ran for several hours, but then died with the error > -similarity_threshold 0.05 -append > decoy_generation.log & > phl004_canonical_sall_osw_decoys_2.1.0.TraML -exclude_similar
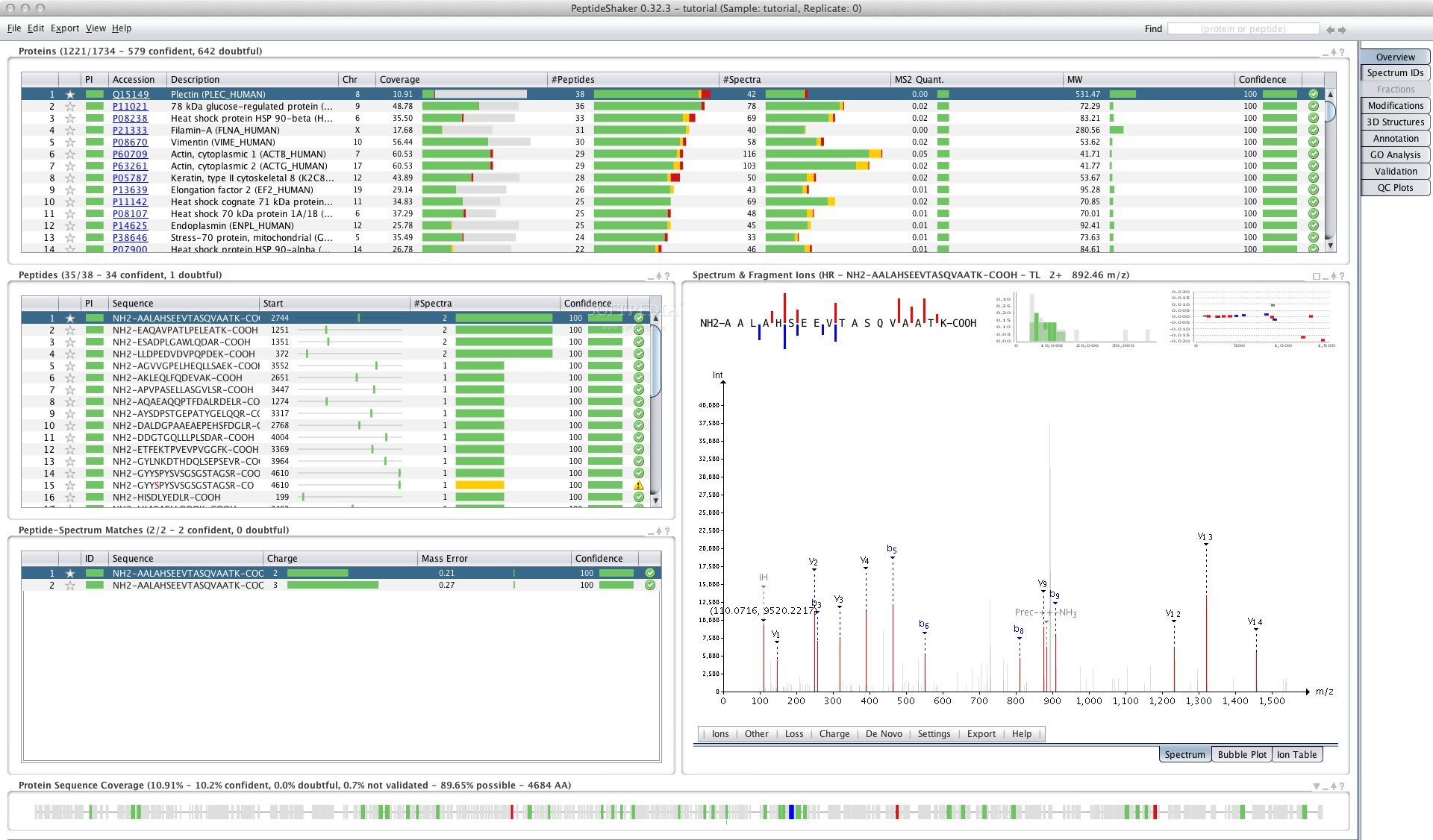
> nohup OpenSwathDecoyGenerator -in phl004_canonical_sall_osw.TraML -out > following command using the new 2.1.0 OpenMS: > did want to be able to generate DECOYS successfully, so I tried the > erroneous mods, I can simply delete them for my current purposes. > "ConvertTraMLToTSV" just to make sure that the input is fine and the > TraML itself is fine, can you run the TraML through > I hope George can comment here as well. > I confess I did have a problem at first when I tried to name the file > So by these metrics the input TraML is correct. > Writing phl004_canonical_sall_osw_traml2tsv.csv > Progress of 'converting to OpenSWATH transition TSV format': > Reading phl004_canonical_sall_osw.TraML > phl004_canonical_sall_osw_traml2tsv.csv > ConvertTraMLToTSV -in phl004_canonical_sall_osw.TraML -out > ConvertTraMLToTSV - Converts a TraML file to an OpenSWATH transition > I'd already checked the xml was well-formed with xmllint -stream, but > data structure to store transitions and OpenSWATH results. > tool finally uses the proper TSV file ending. > ConvertTraMLToTSV are going to be replaced by TargetedFileConverter. > Second, starting with the upcoming OpenMS 2.2, ConvertTSVToTraML and > represent these suggestions in the future. We might also modify the default parameters to > produces a functional TraML that should be directly compatible with the > With the PHL (reconverted from the TSV using ConvertTSVToTraML), this > -remove_unannotated -remove_CNterm_mods > phl004_canonical_s32_osw_decoys.TraML -append -exclude_similar > OpenSwathDecoyGenerator -in phl004_canonical_s32_osw.TraML -out > First, I would recommend to run OpenDecoyGenerator >2.1 with the following > have made some changes that will appear in future OpenMS versions. > I’m sorry to hear about the trouble with the assay generation tools.

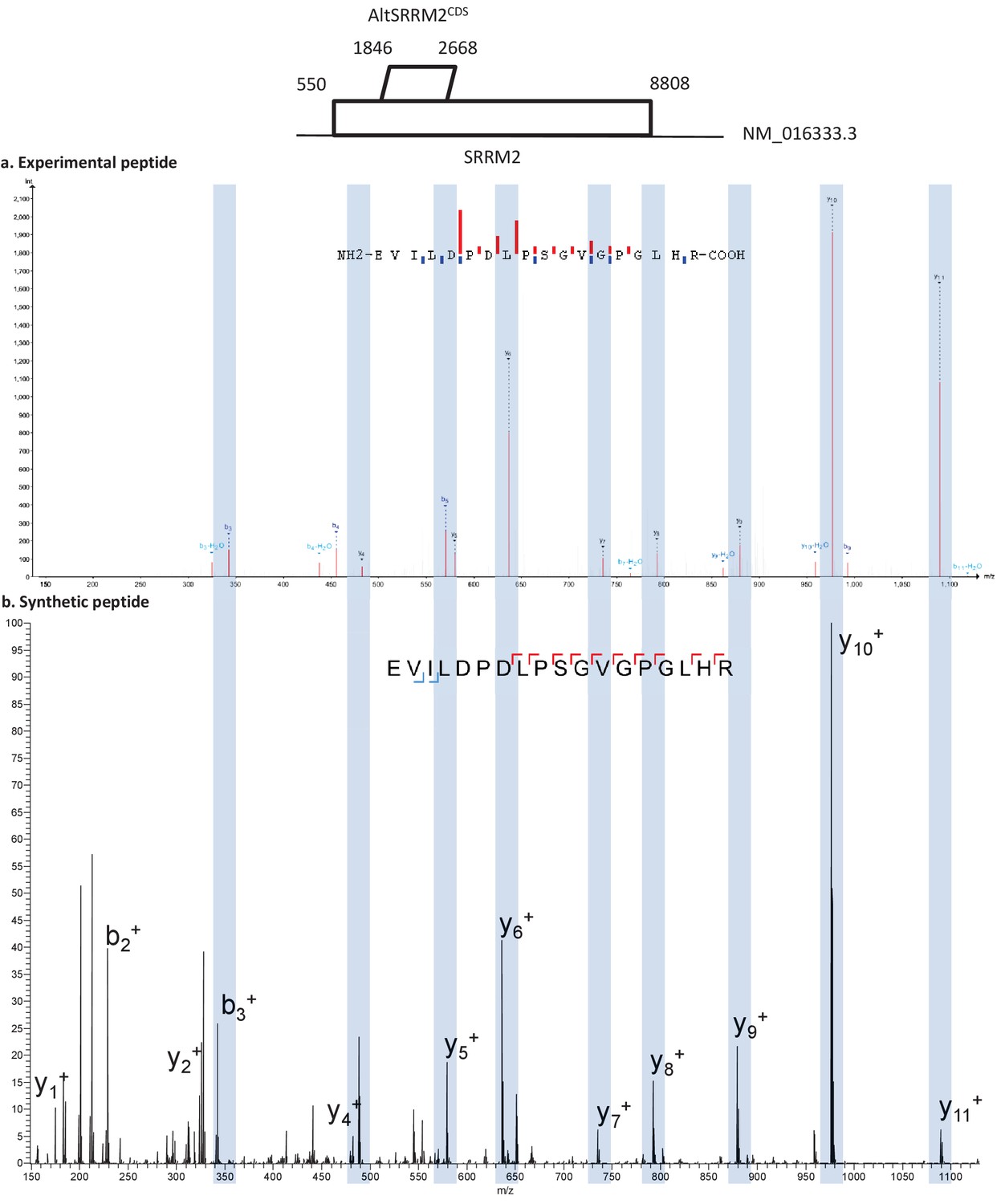
csv files directly, rather than the round-trip through TraML? > Have you considered creating a DECOY generation tool that works on the > Thanks for the detailed response, I'll try with the options you specify. Phl004_canonical_sall_osw_decoys_2.1.0.TraML -exclude_similar OpenSwathDecoyGenerator -in phl004_canonical_s64_ -out TargetedFileConverter -in phl004_canonical_s64_osw.tsv -out Solve this issue is to start with the CSV file: So I have found the currently best way to Now location = 10 indicates that this is a C-terminal modification With newer OpenMS versions and the issue is with existing peptides in I have seen some things the in the TraML that definitely wont work I’m happy to share my input/output files and workflows if this is helpful. If I extend the workflow to do feature identification and peptide quantification, then the featureXML file also doesn’t open - the features are selected (i.e., FeatureFinderCentroided seems to have worked), but features aren’t annotated with the peptide identifications. However, when I use “annotate with identification” in TOPPview, the file doesn’t open, and I can’t see any peptides in the identification view. The MSGF+ search executes properly, and when I view the idXML file either directly as the MSGF+ output or after doing FDR filtering, using TextExporter and IDTextReader, the table looks normal to me. I acquired data on a Thermo Orbitrap Elite, and then converted the files using ProteoWizard to mzML. I’m trying to perform a basic peptide identification and quantification workflow using MSGF+ as the search engine (essentially Figures 7, 10, 11 of the tutorial PDF), except the OMSSAAdapter has been replaced with MSGF+, and I’m running a nonspecific search because I’m looking for endogenous peptides.
